- C Data Types
- C Operators
- C Input and Output
- C Control Flow
- C Functions
- C Preprocessors
- C File Handling
- C Cheatsheet
- C Interview Questions

C Structures
The structure in C is a user-defined data type that can be used to group items of possibly different types into a single type. The struct keyword is used to define the structure in the C programming language. The items in the structure are called its member and they can be of any valid data type. Additionally, the values of a structure are stored in contiguous memory locations.
C Structure Declaration
We have to declare structure in C before using it in our program. In structure declaration, we specify its member variables along with their datatype. We can use the struct keyword to declare the structure in C using the following syntax:
The above syntax is also called a structure template or structure prototype and no memory is allocated to the structure in the declaration. To understand how structures are foundational to building complex data structures, the C Programming Course Online with Data Structures provides practical applications and detailed explanations.
C Structure Definition
To use structure in our program, we have to define its instance. We can do that by creating variables of the structure type. We can define structure variables using two methods:
1. Structure Variable Declaration with Structure Template
2. structure variable declaration after structure template, access structure members.
We can access structure members by using the ( . ) dot operator.
In the case where we have a pointer to the structure, we can also use the arrow operator to access the members.
Initialize Structure Members
Structure members cannot be initialized with the declaration. For example, the following C program fails in the compilation.
The reason for the above error is simple. When a datatype is declared, no memory is allocated for it. Memory is allocated only when variables are created.
Default Initialization
By default, structure members are not automatically initialized to 0 or NULL. Uninitialized structure members will contain garbage values. However, when a structure variable is declared with an initializer, all members not explicitly initialized are zero-initialized.
We can initialize structure members in 3 ways which are as follows:
- Using Assignment Operator.
- Using Initializer List.
- Using Designated Initializer List.
1. Initialization using Assignment Operator
2. initialization using initializer list.
In this type of initialization, the values are assigned in sequential order as they are declared in the structure template.
3. Initialization using Designated Initializer List
Designated Initialization allows structure members to be initialized in any order. This feature has been added in the C99 standard .
The Designated Initialization is only supported in C but not in C++.
Example of Structure in C
The following C program shows how to use structures
typedef for Structures
The typedef keyword is used to define an alias for the already existing datatype. In structures, we have to use the struct keyword along with the structure name to define the variables. Sometimes, this increases the length and complexity of the code. We can use the typedef to define some new shorter name for the structure.
Nested Structures
C language allows us to insert one structure into another as a member. This process is called nesting and such structures are called nested structures . There are two ways in which we can nest one structure into another:
1. Embedded Structure Nesting
In this method, the structure being nested is also declared inside the parent structure.
2. Separate Structure Nesting
In this method, two structures are declared separately and then the member structure is nested inside the parent structure.
One thing to note here is that the declaration of the structure should always be present before its definition as a structure member. For example, the declaration below is invalid as the struct mem is not defined when it is declared inside the parent structure.
Accessing Nested Members
We can access nested Members by using the same ( . ) dot operator two times as shown:
Example of Structure Nesting
Structure pointer in c.
We can define a pointer that points to the structure like any other variable. Such pointers are generally called Structure Pointers . We can access the members of the structure pointed by the structure pointer using the ( -> ) arrow operator.
Example of Structure Pointer
Self-referential structures.
The self-referential structures in C are those structures that contain references to the same type as themselves i.e. they contain a member of the type pointer pointing to the same structure type.
Example of Self-Referential Structures
Such kinds of structures are used in different data structures such as to define the nodes of linked lists, trees, etc.
C Structure Padding and Packing
Technically, the size of the structure in C should be the sum of the sizes of its members. But it may not be true for most cases. The reason for this is Structure Padding.
Structure padding is the concept of adding multiple empty bytes in the structure to naturally align the data members in the memory. It is done to minimize the CPU read cycles to retrieve different data members in the structure.
There are some situations where we need to pack the structure tightly by removing the empty bytes. In such cases, we use Structure Packing. C language provides two ways for structure packing:
- Using #pragma pack(1)
- Using __attribute((packed))__
Example of Structure Padding and Packing
As we can see, the size of the structure is varied when structure packing is performed.
To know more about structure padding and packing, refer to this article – Structure Member Alignment, Padding and Data Packing .
Bit Fields are used to specify the length of the structure members in bits. When we know the maximum length of the member, we can use bit fields to specify the size and reduce memory consumption.
Syntax of Bit Fields
example of bit fields.
As we can see, the size of the structure is reduced when using the bit field to define the max size of the member ‘a’.
Uses of Structure in C
C structures are used for the following:
- The structure can be used to define the custom data types that can be used to create some complex data types such as dates, time, complex numbers, etc. which are not present in the language.
- It can also be used in data organization where a large amount of data can be stored in different fields.
- Structures are used to create data structures such as trees, linked lists, etc.
- They can also be used for returning multiple values from a function.
Limitations of C Structures
In C language, structures provide a method for packing together data of different types. A Structure is a helpful tool to handle a group of logically related data items. However, C structures also have some limitations.
- Higher Memory Consumption: It is due to structure padding.
- No Data Hiding: C Structures do not permit data hiding. Structure members can be accessed by any function, anywhere in the scope of the structure.
- Functions inside Structure: C structures do not permit functions inside the structure so we cannot provide the associated functions.
- Static Members: C Structure cannot have static members inside its body.
- Construction creation in Structure: Structures in C cannot have a constructor inside Structures.
Related Articles
- C Structures vs C++ Structure
Similar Reads
- C-Structure & Union
Improve your Coding Skills with Practice
What kind of Experience do you want to share?
Next: Unions , Previous: Overlaying Structures , Up: Structures [ Contents ][ Index ]
15.13 Structure Assignment
Assignment operating on a structure type copies the structure. The left and right operands must have the same type. Here is an example:
Notionally, assignment on a structure type works by copying each of the fields. Thus, if any of the fields has the const qualifier, that structure type does not allow assignment:
See Assignment Expressions .
When a structure type has a field which is an array, as here,
structure assigment such as r1 = r2 copies array fields’ contents just as it copies all the other fields.
This is the only way in C that you can operate on the whole contents of a array with one operation: when the array is contained in a struct . You can’t copy the contents of the data field as an array, because
would convert the array objects (as always) to pointers to the zeroth elements of the arrays (of type struct record * ), and the assignment would be invalid because the left operand is not an lvalue.
C Functions
C structures, c reference, c structures (structs).
Structures (also called structs) are a way to group several related variables into one place. Each variable in the structure is known as a member of the structure.
Unlike an array , a structure can contain many different data types (int, float, char, etc.).
Create a Structure
You can create a structure by using the struct keyword and declare each of its members inside curly braces:
To access the structure, you must create a variable of it.
Use the struct keyword inside the main() method, followed by the name of the structure and then the name of the structure variable:
Create a struct variable with the name "s1":
Access Structure Members
To access members of a structure, use the dot syntax ( . ):
Now you can easily create multiple structure variables with different values, using just one structure:
Advertisement
What About Strings in Structures?
Remember that strings in C are actually an array of characters, and unfortunately, you can't assign a value to an array like this:
An error will occur:
However, there is a solution for this! You can use the strcpy() function and assign the value to s1.myString , like this:
Simpler Syntax
You can also assign values to members of a structure variable at declaration time, in a single line.
Just insert the values in a comma-separated list inside curly braces {} . Note that you don't have to use the strcpy() function for string values with this technique:
Note: The order of the inserted values must match the order of the variable types declared in the structure (13 for int, 'B' for char, etc).
Copy Structures
You can also assign one structure to another.
In the following example, the values of s1 are copied to s2:
Modify Values
If you want to change/modify a value, you can use the dot syntax ( . ).
And to modify a string value, the strcpy() function is useful again:
Modifying values are especially useful when you copy structure values:
Ok, so, how are structures useful?
Imagine you have to write a program to store different information about Cars, such as brand, model, and year. What's great about structures is that you can create a single "Car template" and use it for every cars you make. See below for a real life example.
Real-Life Example
Use a structure to store different information about Cars:

COLOR PICKER

Contact Sales
If you want to use W3Schools services as an educational institution, team or enterprise, send us an e-mail: [email protected]
Report Error
If you want to report an error, or if you want to make a suggestion, send us an e-mail: [email protected]
Top Tutorials
Top references, top examples, get certified.
Learn C practically and Get Certified .

Popular Tutorials
Popular examples, reference materials, certification courses.
Created with over a decade of experience and thousands of feedback.
C Introduction
- Getting Started with C
- Your First C Program
C Fundamentals
- C Variables, Constants and Literals
- C Data Types
- C Input Output (I/O)
- C Programming Operators
C Flow Control
- C if...else Statement
- C while and do...while Loop
- C break and continue
- C switch Statement
- C goto Statement
- C Functions
- C User-defined functions
- Types of User-defined Functions in C Programming
- C Recursion
- C Storage Class
C Programming Arrays
- C Multidimensional Arrays
- Pass arrays to a function in C
C Programming Pointers
- Relationship Between Arrays and Pointers
- C Pass Addresses and Pointers
- C Dynamic Memory Allocation
- C Array and Pointer Examples
- C Programming Strings
- String Manipulations In C Programming Using Library Functions
- String Examples in C Programming
C Structure and Union
C structs and Pointers
C Structure and Function
C Programming Files
- C File Handling
- C Files Examples
C Additional Topics
- C Keywords and Identifiers
- C Precedence And Associativity Of Operators
- C Bitwise Operators
- C Preprocessor and Macros
- C Standard Library Functions
C Tutorials
C Struct Examples
- Add Two Complex Numbers by Passing Structure to a Function
- Add Two Distances (in inch-feet system) using Structures
In C programming, a struct (or structure) is a collection of variables (can be of different types) under a single name.
- Define Structures
Before you can create structure variables, you need to define its data type. To define a struct, the struct keyword is used.
Syntax of struct
For example,
Here, a derived type struct Person is defined. Now, you can create variables of this type.
Create struct Variables
When a struct type is declared, no storage or memory is allocated. To allocate memory of a given structure type and work with it, we need to create variables.
Here's how we create structure variables:
Another way of creating a struct variable is:
In both cases,
- person1 and person2 are struct Person variables
- p[] is a struct Person array of size 20.
Access Members of a Structure
There are two types of operators used for accessing members of a structure.
- . - Member operator
- -> - Structure pointer operator (will be discussed in the next tutorial)
Suppose, you want to access the salary of person2 . Here's how you can do it.
Example 1: C structs
In this program, we have created a struct named Person . We have also created a variable of Person named person1 .
In main() , we have assigned values to the variables defined in Person for the person1 object.
Notice that we have used strcpy() function to assign the value to person1.name .
This is because name is a char array ( C-string ) and we cannot use the assignment operator = with it after we have declared the string.
Finally, we printed the data of person1 .
- Keyword typedef
We use the typedef keyword to create an alias name for data types. It is commonly used with structures to simplify the syntax of declaring variables.
For example, let us look at the following code:
We can use typedef to write an equivalent code with a simplified syntax:
Example 2: C typedef
Here, we have used typedef with the Person structure to create an alias person .
Now, we can simply declare a Person variable using the person alias:
- Nested Structures
You can create structures within a structure in C programming. For example,
Suppose, you want to set imag of num2 variable to 11 . Here's how you can do it:
Example 3: C Nested Structures
Why structs in c.
Suppose you want to store information about a person: his/her name, citizenship number, and salary. You can create different variables name , citNo and salary to store this information.
What if you need to store information of more than one person? Now, you need to create different variables for each information per person: name1 , citNo1 , salary1 , name2 , citNo2 , salary2 , etc.
A better approach would be to have a collection of all related information under a single name Person structure and use it for every person.
More on struct
- Structures and pointers
- Passing structures to a function
Table of Contents
- C struct (Introduction)
- Create struct variables
- Access members of a structure
- Example 1: C++ structs
Sorry about that.
Our premium learning platform, created with over a decade of experience and thousands of feedbacks .
Learn and improve your coding skills like never before.
- Interactive Courses
- Certificates
- 2000+ Challenges
Related Tutorials

Structures in C programming, need and use
- Need of structures?
- What is structure?
- How to declare and define?
- How to create structure object?
- How to access structure members?
- Example program
Structures in C, is an advance and most popular topic in C language. It facilitates you to design your custom data type . In this tutorial, we will learn about structures in C its need, how to declare, define and access structures.
Need of structures in C?
C has built in primitive and derrived data types . Still not all real world problems can be solved using those types. You need custom data type for different situations.
For example, if you need to store 100 student record that consist of name, age and mobile number. To code that you will create 3 array variables each of size 100 i.e. name[100] , age[100] , mobile[100] . For three fields in student record it say seem feasible to you. But, think how cumbersome it would be to manage student record with more than 10 fields, in separate variables for single student.
To overcome this we need a user defined data type. In this tutorial I am going to explain how easily we will deal with these situations using structures in C programming language.
What is structure in C?
Structure is a user defined data type. It is a collection of different data type, to create a new data type.
For example, You can define your custom type for storing student record containing name, age and mobile. Creating structure type will allow you to handle all properties (fields) of student with single variable, instead of handling it separately.
How to declare, define and access structure members?
To declare or define a structure, we use struct keyword. It is a reserved word in the C compiler. You must only use it for structure or its object declaration or definition.
Syntax to define a structure
Here, structure_name is name of our custom type. memberN_declaration is structure member i.e. variable declaration that structure will have.
Example to define a structure
Let us use our student example and define a structure to store student object.
Points to remember while structure definition
- You must terminate structure definition with semicolon ; .
For example, following is an invalid structure definition.
- You can define a structure anywhere like global scope (accessible by all functions) or local scope (accessible by particular function).
- Structure member definition may contain other structure type.
How to create structure object (structure variable)?
A data type is useless without variables. A data type defines various properties about data stored in memory. To use any type we must declare its variable. Hence, let us learn how to create our custom structure type objects also known as structure variable .
In C programming, there are two ways to declare a structure variable:
- Along with structure definition
- After structure definition
Declaration along with the structure definition
Out of two ways to declare structure variable. You can declare a structure variable along with structure before terminating the structure definition.
So, if you want to declare student type object along with student structure definition you can use this approach.
Declaration after structure definition
The other way to declare, gives you luxury to declare structure variable anywhere in program based on the structure scope. If structure is defined in global scope, we can declare its variable in main() function , any other functions and in the global section too.
For above example, if we want to declare its variable with name student1 , it will be declared as given below:
How to access structure members (data)?
You created structure and its variable. But since structure is a complex data type, you cannot assign any value directly to it using assignment operator . You must assign data to individual structure members separately.
C supports two operators to access structure members, using a structure variable.
- Dot/period operator .
- Arrow operator ->
Dot/period operator (.) in C
Dot/period operator also known as member access operator. We use dot operator to access members of simple structure variable.
Read more about other operators in C programming language .
Arrow operator (->) in C
Since structure is a user defined type and you can have pointers to any type. Hence, you may also create pointers to structure.
In C language it is illegal to access a structure member from a pointer to structure variable using dot operator. We use arrow operator -> to access structure member from pointer to structure.
Example program to demonstrate declare, define and access structure members
In this example, we will declare a structure type, create structure object and access structure members. I will show how to use both dot and arrow operator to access structure members.
Happy coding 😉
- C Programming
- C Basic Syntax
- C Data Types
- C if...else Statement
- C switch Statement
- C while Loop
- C do...while Loop
- C Jump Statements
- C Functions
- C Library Functions
- C Recursion
- C Variable Scope
- C Structures
- C Linked List
- C Binary Tree
- C Header Files
- C Programming Examples
Structures in C programming with examples
It is possible to combine data items of various types using the structure, which is a user-defined data type. In other words, the structure is nothing more than a collection of variables that are grouped together under a single name, which is referred to as a "aggregate data type."
Define a Structure in C
To define a structure in C, all you have to do is use the "struct" keyword. The following is the general form that must be used when defining a structure in C:
Here "struct" is a keyword used in declaring a structure in C. Then struct_name is the name of the structure. Then type can be any valid data type. And then, member_name1, member_name2, member_name3, and member_nameN are the names of the structure members. Finally, structure_variable_name is the name of the structure variable that is used to access a member of this structure using the dot operator. You can also declare structure_variable_name later. You can also use this general form.
Let's look at this declaration to understand how to define a structure in the C language:
Here we define a structure type called address and declare a variable named addr_var of that type. When a structure variable (addr_var here) is declared, then the compiler automatically allocates sufficient memory to accommodate all of its members.
Access Structure Members in C
To access individual members of a structure in C, simply use the dot (.) operator. Here is the general form to access structure members in C:
Let's look at the following code fragment, demonstrating how to access structure members:
And to print the ZIP code on the screen, follow this statement:
The above statement shows the zip code from the structure variable addr_var's zip member.
Structure Assignments in C
In C, one structure can also be assigned to another structure of the same type. Using a single assignment statement, the information contained in one structure can be assigned to another structure of the same type. The following is the standard format for assigning one structure to another of the same type:
Here is an example program demonstrating structure assignment:
The following snapshot shows the initial output produced by the above program:
Now enter your name, "William," and press the ENTER key. Then enter three numbers in the following order: enter the first number, say 10, and press the ENTER key, then enter the second number, say 20, and press the ENTER key, and finally type the third number, say 30, and press the ENTER key to get the following output:
Now, type "y" and press the ENTER key to see the sum of the three numbers you entered on the output console, as shown in the screenshot below:
As you can see, this program gets the value entered by the user and stores it in the structure members accessed by the first structure (the structure variable), and then we assign it to another structure (the structure variable). So that all the members' values in the first structure can be accessed using the second structure (structure variables) as well.
The method "fflush()" is used to flush the output buffer.
Arrays of Structures in C
Typically, structures are arrayed. To declare an array of structures, you must first define a structure, followed by the declaration of an array variable of that type. Here is an instance of declaring a 10-element structure array of type st:
The preceding statement generates ten sets of variables organized in accordance with the st structure.
To gain access to a particular structure, simply index the array's name. Here is an example of accessing the structure 4 member zip:
Indexing always begins with 0, so index number 0 refers to the first element, index number 1 to the second element, and index number 3 to the fourth element. Here's an example of how to assign a value to structure 4's member zip:
Here is an example of printing the value of member zip in structure 4:
Here is an example, assigning "A" to the first character of name in the fourth structure of st_var:
Let's look at the following example for a complete understanding of the concept of arrays of structures in the C language:
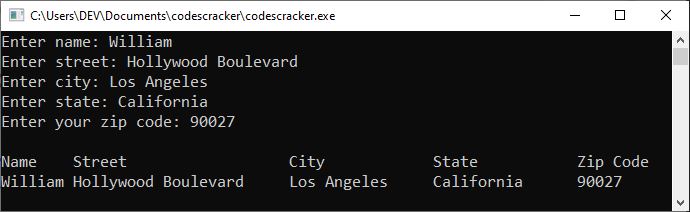
The following snapshot shows the sample run after providing the following inputs:
- "William" as the name.
- "Hollywood Boulevard" as the street.
- "Los Angeles" as the city.
- "California" as the state.
- "90027" as the zip code.
Pass structures to functions in C
In C, it is permissible to pass structure members or entire structures to functions. Whenever you pass a structure member to a function, you are passing the member's value to the function. Consider the following structure declaration:
Here is an example of passing each member to a function:
Here is another example of passing the address of each member to a function:
When a structure is passed as an argument to a function in this instance, the entire structure is passed using the standard call-by-value method. Here is an example of passing an entire structure to a function:
The following snapshot shows the sample run.
Structure Pointers in C
In the C programming language, pointers to structures are permitted. Here is the general form of a structure pointer declaration in C:
To determine the address of any structure variable, prefix the structure's name with the & operator. Consider the following structure declaration:
Now the following code fragment simply places the address of the structure stc_var into the pointer ptr:
And you must use the "->" operator to access the members of a structure using a pointer to that structure. Here is an illustration using the "num" field:
Here is an illustration of structure pointers in the C programming language. Before I include the program, I'd like to point out that it continues to print the time.
Here is a sample run of this C program. This program continues to display the time starting from 00:00:00.
C Online Test
Liked this post? Share it!

IMAGES
VIDEO
COMMENTS
Yes, you can assign one instance of a struct to another using a simple assignment statement. In the case of non-pointer or non pointer containing struct members, assignment means copy. In the case of pointer struct members, assignment means pointer will point to the same address of the other pointer. Let us see this first hand:
The structure in C is a user-defined data type that can be used to group items of possibly different types into a single type. The struct keyword is used to define the structure in the C programming language. The items in the structure are called its member and they can be of any valid data type. Additionally, the values of a structure are stored in contiguous memory locations.
Example of Nested Structure in C Programming. Let's say we have two structure like this: The second structure stu_data has stu_address as a data member. Here, stu_data is called outer structure or parent structure and stu_address is called inner structure or child structure. Structure 1: stu_address
structure assigment such as r1 = r2 copies array fields' contents just as it copies all the other fields.. This is the only way in C that you can operate on the whole contents of a array with one operation: when the array is contained in a struct.You can't copy the contents of the data field as an array, because
C Structures C Structures C Enums C Enums C Memory C Memory Management. ... prog.c:12:15: error: assignment to expression with array type. Try it Yourself » ... Imagine you have to write a program to store different information about Cars, such as brand, model, and year. What's great about structures is that you can create a single "Car ...
@gabeappleton Struct assignment always replace left side struct with the data from right side. On the right side of your sample is structure with designated initializer which missing fields will be initialized to zero by default. And then s1 will be replaced with content of new structure. So id field will be set to zero. -
A Structure is a collection of related data items, possibly of different types. Structures are also called records. A structure type in C is called struct. Unlike arrays, a struct is composed of data of different types. You use structures to group data that belong together. Examples: o Student information: student id,
In this tutorial, you'll learn about struct types in C Programming. You will learn to define and use structures with the help of examples. In C programming, a struct (or structure) is a collection of variables (can be of different types) under a single name. ... and we cannot use the assignment operator = with it after we have declared the ...
Hence, let us learn how to create our custom structure type objects also known as structure variable. In C programming, there are two ways to declare a structure variable: ... you cannot assign any value directly to it using assignment operator. You must assign data to individual structure members separately. C supports two operators to access ...
Structures in C programming with examples: The structure is nothing more than a collection of variables that are grouped together under a single name, which is referred to as a aggregate data type. ... Structure Assignments in C. In C, one structure can also be assigned to another structure of the same type. Using a single assignment statement ...